Rows: 1,236
Columns: 8
$ case <int> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 1…
$ bwt <int> 120, 113, 128, 123, 108, 136, 138, 132, 120, 143, 140, 144, …
$ gestation <int> 284, 282, 279, NA, 282, 286, 244, 245, 289, 299, 351, 282, 2…
$ parity <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, …
$ age <int> 27, 33, 28, 36, 23, 25, 33, 23, 25, 30, 27, 32, 23, 36, 30, …
$ height <int> 62, 64, 64, 69, 67, 62, 62, 65, 62, 66, 68, 64, 63, 61, 63, …
$ weight <int> 100, 135, 115, 190, 125, 93, 178, 140, 125, 136, 120, 124, 1…
$ smoke <int> 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 0, 1, 1, 1, 0, 0, 1, 1, 0, 1, …Multiple Linear Regression, Transformations, and Model Evaluation
Data babies in openintro package
| y | Response | Birth weight | Numeric |
|---|---|---|---|
| \(x_1\) | Explanatory | Gestation | Numeric |
| \(x_2\) | Explanatory | Smoke | Categorical |
Notation
\(y_i = \beta_0 +\beta_1x_{1i} + \beta_2x_{2i} + \epsilon_i\)
\(\beta_0\) is intercept
\(\beta_1\) is the slope for gestation
\(\beta_2\) is the slope for smoke
\(\epsilon_i\) is error/residual
\(i = 1, 2, ...n\) identifier for each point
Expected birth weight for a baby who had 280 days of gestation with a smoker mother
\(\hat {\text{bwt}_i} = b_0 + b_1 \text{ gestation}_i + b_2 \text{ smoke}_i\)
\(\hat {\text{bwt}_i} = -0.932 + (0.443 \times 280) + (-8.09 \times 1)\)
# A tibble: 1,236 × 4
bwt gestation smoke pred
<int> <int> <int> <dbl>
1 120 284 0 125.
2 113 282 0 124.
3 128 279 1 115.
4 123 NA 0 NA
5 108 282 1 116.
6 136 286 0 126.
7 138 244 0 107.
8 132 245 0 108.
9 120 289 0 127.
10 143 299 1 123.
# ℹ 1,226 more rows# A tibble: 1,236 × 5
bwt gestation smoke pred resid
<int> <int> <int> <dbl> <dbl>
1 120 284 0 125. -4.84
2 113 282 0 124. -11.0
3 128 279 1 115. 13.5
4 123 NA 0 NA NA
5 108 282 1 116. -7.87
6 136 286 0 126. 10.3
7 138 244 0 107. 30.9
8 132 245 0 108. 24.4
9 120 289 0 127. -7.05
10 143 299 1 123. 19.6
# ℹ 1,226 more rowsTransformations
Data
Rows: 2,930
Columns: 82
$ order <int> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16,…
$ pid <chr> "0526301100", "0526350040", "0526351010", "0526353030"…
$ ms_sub_class <chr> "020", "020", "020", "020", "060", "060", "120", "120"…
$ ms_zoning <chr> "RL", "RH", "RL", "RL", "RL", "RL", "RL", "RL", "RL", …
$ lot_frontage <int> 141, 80, 81, 93, 74, 78, 41, 43, 39, 60, 75, NA, 63, 8…
$ lot_area <int> 31770, 11622, 14267, 11160, 13830, 9978, 4920, 5005, 5…
$ street <chr> "Pave", "Pave", "Pave", "Pave", "Pave", "Pave", "Pave"…
$ alley <chr> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
$ lot_shape <chr> "IR1", "Reg", "IR1", "Reg", "IR1", "IR1", "Reg", "IR1"…
$ land_contour <chr> "Lvl", "Lvl", "Lvl", "Lvl", "Lvl", "Lvl", "Lvl", "HLS"…
$ utilities <chr> "AllPub", "AllPub", "AllPub", "AllPub", "AllPub", "All…
$ lot_config <chr> "Corner", "Inside", "Corner", "Corner", "Inside", "Ins…
$ land_slope <chr> "Gtl", "Gtl", "Gtl", "Gtl", "Gtl", "Gtl", "Gtl", "Gtl"…
$ neighborhood <chr> "NAmes", "NAmes", "NAmes", "NAmes", "Gilbert", "Gilber…
$ condition_1 <chr> "Norm", "Feedr", "Norm", "Norm", "Norm", "Norm", "Norm…
$ condition_2 <chr> "Norm", "Norm", "Norm", "Norm", "Norm", "Norm", "Norm"…
$ bldg_type <chr> "1Fam", "1Fam", "1Fam", "1Fam", "1Fam", "1Fam", "Twnhs…
$ house_style <chr> "1Story", "1Story", "1Story", "1Story", "2Story", "2St…
$ overall_qual <int> 6, 5, 6, 7, 5, 6, 8, 8, 8, 7, 6, 6, 6, 7, 8, 8, 8, 9, …
$ overall_cond <int> 5, 6, 6, 5, 5, 6, 5, 5, 5, 5, 5, 7, 5, 5, 5, 5, 7, 2, …
$ year_built <int> 1960, 1961, 1958, 1968, 1997, 1998, 2001, 1992, 1995, …
$ year_remod_add <int> 1960, 1961, 1958, 1968, 1998, 1998, 2001, 1992, 1996, …
$ roof_style <chr> "Hip", "Gable", "Hip", "Hip", "Gable", "Gable", "Gable…
$ roof_matl <chr> "CompShg", "CompShg", "CompShg", "CompShg", "CompShg",…
$ exterior_1st <chr> "BrkFace", "VinylSd", "Wd Sdng", "BrkFace", "VinylSd",…
$ exterior_2nd <chr> "Plywood", "VinylSd", "Wd Sdng", "BrkFace", "VinylSd",…
$ mas_vnr_type <chr> "Stone", "None", "BrkFace", "None", "None", "BrkFace",…
$ mas_vnr_area <int> 112, 0, 108, 0, 0, 20, 0, 0, 0, 0, 0, 0, 0, 0, 0, 603,…
$ exter_qual <chr> "TA", "TA", "TA", "Gd", "TA", "TA", "Gd", "Gd", "Gd", …
$ exter_cond <chr> "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", …
$ foundation <chr> "CBlock", "CBlock", "CBlock", "CBlock", "PConc", "PCon…
$ bsmt_qual <chr> "TA", "TA", "TA", "TA", "Gd", "TA", "Gd", "Gd", "Gd", …
$ bsmt_cond <chr> "Gd", "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", …
$ bsmt_exposure <chr> "Gd", "No", "No", "No", "No", "No", "Mn", "No", "No", …
$ bsmt_fin_type_1 <chr> "BLQ", "Rec", "ALQ", "ALQ", "GLQ", "GLQ", "GLQ", "ALQ"…
$ bsmt_fin_sf_1 <int> 639, 468, 923, 1065, 791, 602, 616, 263, 1180, 0, 0, 9…
$ bsmt_fin_type_2 <chr> "Unf", "LwQ", "Unf", "Unf", "Unf", "Unf", "Unf", "Unf"…
$ bsmt_fin_sf_2 <int> 0, 144, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1120, 0, 0…
$ bsmt_unf_sf <int> 441, 270, 406, 1045, 137, 324, 722, 1017, 415, 994, 76…
$ total_bsmt_sf <int> 1080, 882, 1329, 2110, 928, 926, 1338, 1280, 1595, 994…
$ heating <chr> "GasA", "GasA", "GasA", "GasA", "GasA", "GasA", "GasA"…
$ heating_qc <chr> "Fa", "TA", "TA", "Ex", "Gd", "Ex", "Ex", "Ex", "Ex", …
$ central_air <chr> "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y",…
$ electrical <chr> "SBrkr", "SBrkr", "SBrkr", "SBrkr", "SBrkr", "SBrkr", …
$ x1st_flr_sf <int> 1656, 896, 1329, 2110, 928, 926, 1338, 1280, 1616, 102…
$ x2nd_flr_sf <int> 0, 0, 0, 0, 701, 678, 0, 0, 0, 776, 892, 0, 676, 0, 0,…
$ low_qual_fin_sf <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, …
$ gr_liv_area <int> 1656, 896, 1329, 2110, 1629, 1604, 1338, 1280, 1616, 1…
$ bsmt_full_bath <int> 1, 0, 0, 1, 0, 0, 1, 0, 1, 0, 0, 1, 0, 1, 1, 1, 0, 1, …
$ bsmt_half_bath <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, …
$ full_bath <int> 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 1, 1, 3, 2, 1, …
$ half_bath <int> 0, 0, 1, 1, 1, 1, 0, 0, 0, 1, 1, 0, 1, 1, 1, 1, 0, 1, …
$ bedroom_abv_gr <int> 3, 2, 3, 3, 3, 3, 2, 2, 2, 3, 3, 3, 3, 2, 1, 4, 4, 1, …
$ kitchen_abv_gr <int> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
$ kitchen_qual <chr> "TA", "TA", "Gd", "Ex", "TA", "Gd", "Gd", "Gd", "Gd", …
$ tot_rms_abv_grd <int> 7, 5, 6, 8, 6, 7, 6, 5, 5, 7, 7, 6, 7, 5, 4, 12, 8, 8,…
$ functional <chr> "Typ", "Typ", "Typ", "Typ", "Typ", "Typ", "Typ", "Typ"…
$ fireplaces <int> 2, 0, 0, 2, 1, 1, 0, 0, 1, 1, 1, 0, 1, 1, 0, 1, 0, 1, …
$ fireplace_qu <chr> "Gd", NA, NA, "TA", "TA", "Gd", NA, NA, "TA", "TA", "T…
$ garage_type <chr> "Attchd", "Attchd", "Attchd", "Attchd", "Attchd", "Att…
$ garage_yr_blt <int> 1960, 1961, 1958, 1968, 1997, 1998, 2001, 1992, 1995, …
$ garage_finish <chr> "Fin", "Unf", "Unf", "Fin", "Fin", "Fin", "Fin", "RFn"…
$ garage_cars <int> 2, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 3, 2, 3, …
$ garage_area <int> 528, 730, 312, 522, 482, 470, 582, 506, 608, 442, 440,…
$ garage_qual <chr> "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", …
$ garage_cond <chr> "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", "TA", …
$ paved_drive <chr> "P", "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y", "Y",…
$ wood_deck_sf <int> 210, 140, 393, 0, 212, 360, 0, 0, 237, 140, 157, 483, …
$ open_porch_sf <int> 62, 0, 36, 0, 34, 36, 0, 82, 152, 60, 84, 21, 75, 0, 5…
$ enclosed_porch <int> 0, 0, 0, 0, 0, 0, 170, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0…
$ x3ssn_porch <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, …
$ screen_porch <int> 0, 120, 0, 0, 0, 0, 0, 144, 0, 0, 0, 0, 0, 0, 140, 210…
$ pool_area <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, …
$ pool_qc <chr> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
$ fence <chr> NA, "MnPrv", NA, NA, "MnPrv", NA, NA, NA, NA, NA, NA, …
$ misc_feature <chr> NA, NA, "Gar2", NA, NA, NA, NA, NA, NA, NA, NA, "Shed"…
$ misc_val <int> 0, 0, 12500, 0, 0, 0, 0, 0, 0, 0, 0, 500, 0, 0, 0, 0, …
$ mo_sold <int> 5, 6, 6, 4, 3, 6, 4, 1, 3, 6, 4, 3, 5, 2, 6, 6, 6, 6, …
$ yr_sold <int> 2010, 2010, 2010, 2010, 2010, 2010, 2010, 2010, 2010, …
$ sale_type <chr> "WD ", "WD ", "WD ", "WD ", "WD ", "WD ", "WD ", "WD "…
$ sale_condition <chr> "Normal", "Normal", "Normal", "Normal", "Normal", "Nor…
$ sale_price <int> 215000, 105000, 172000, 244000, 189900, 195500, 213500…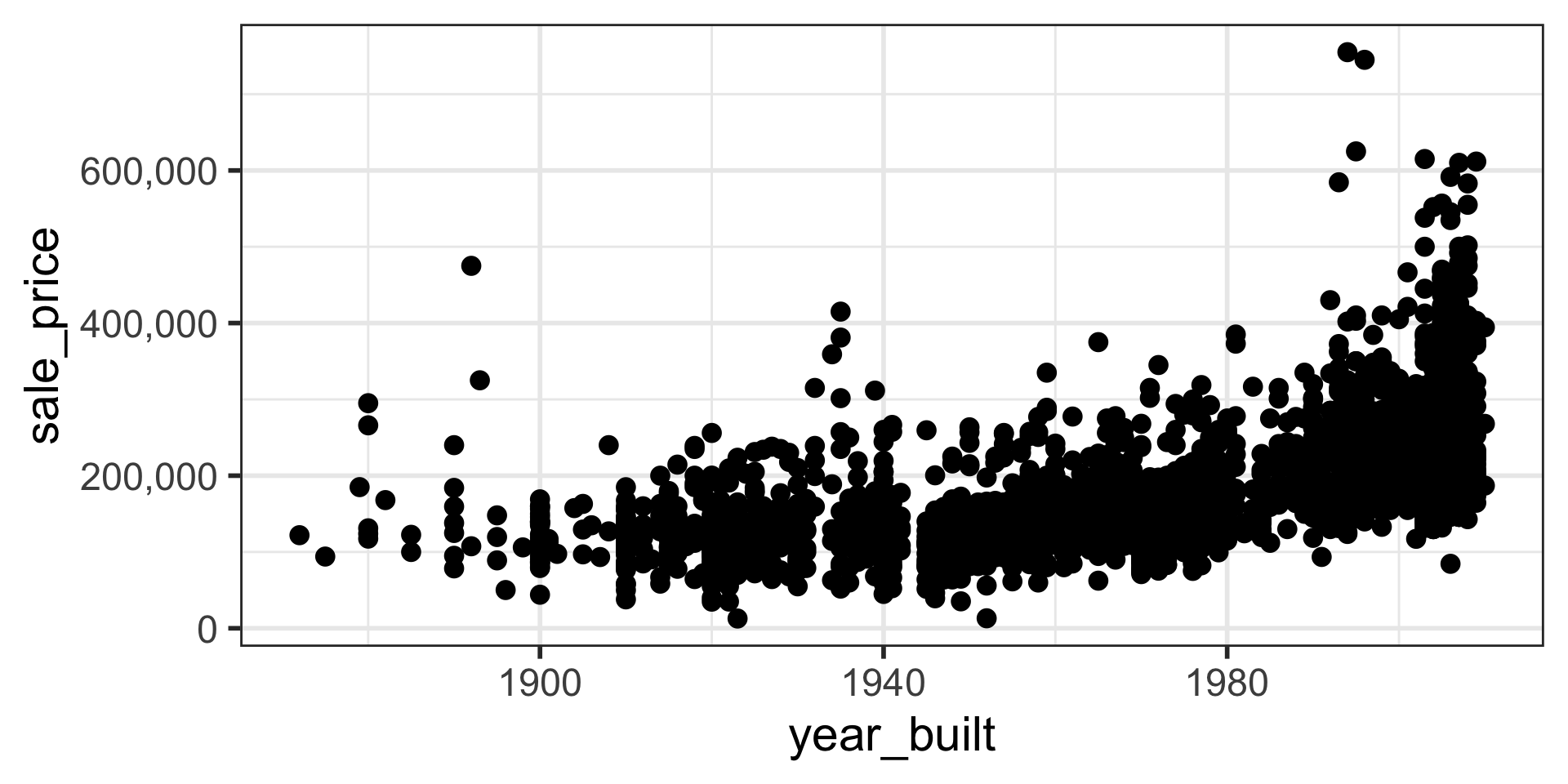
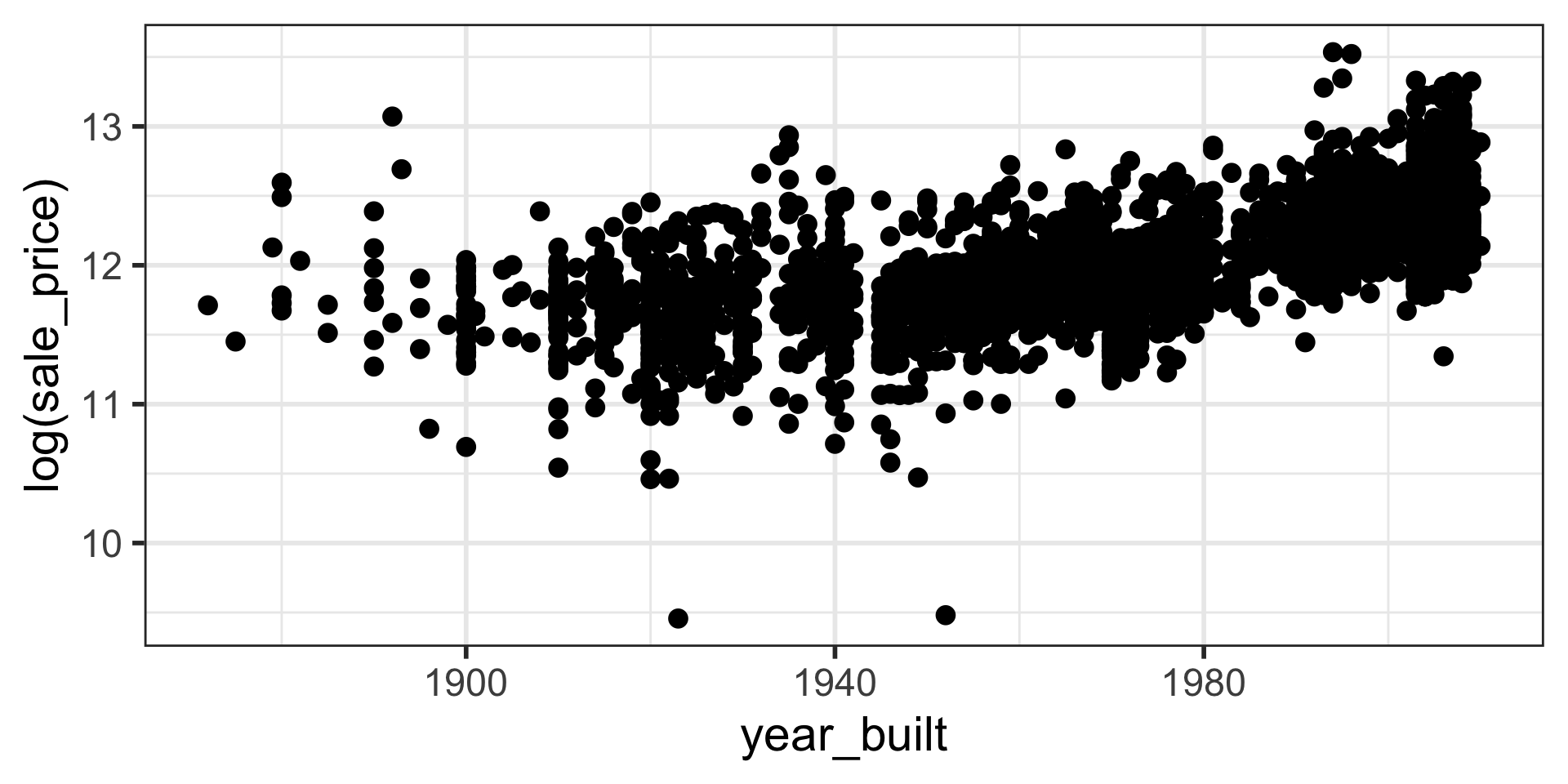
Note that log is natural log in R.
# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -4.33 0.387 -11.2 1.73e- 28
2 year_built 0.00829 0.000196 42.3 4.45e-305\({log(\hat y_i)} = b_0 + b_1x_{1i}\)
\({log(\hat y_i)} = -4.33 + 0.00829x_{1i}\)
Estimated sale price of a house built in 1980
\({log(\hat y_i)} = -4.33 + 0.00829 \times 1980\)
\(e^{log(\hat y_i)} = e^{-4.33 + 0.00829 \times 1980}\)
\(\hat y_i = e^{-4.33} \times e^ {0.00829 \times 1980} = 177052.2\)
For one-unit (year) increase in x, the y is multiplied by \(e^{b_1}\).
Note that \(e^x\) can be achieved by exp(x) in R.
When Relationships Aren’t Linear: Transformations
- Linear regression models assume a linear relationship between the explanatory (X) and response (Y) variables.
- What if the real relationship is curved (non-linear)?
- We can transform our variables to make the relationship linear, allowing us to still use linear regression.
Common Transformations
- Logarithmic:
log(Y)orlog(X). Useful when a variable has a wide range of values or a skewed distribution (like housing prices). - Reciprocal:
1/Yor1/X - Square Root:
sqrt(Y)orsqrt(X)
Model Evaluation
Rows: 1,236
Columns: 8
$ case <int> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 1…
$ bwt <int> 120, 113, 128, 123, 108, 136, 138, 132, 120, 143, 140, 144, …
$ gestation <int> 284, 282, 279, NA, 282, 286, 244, 245, 289, 299, 351, 282, 2…
$ parity <int> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, …
$ age <int> 27, 33, 28, 36, 23, 25, 33, 23, 25, 30, 27, 32, 23, 36, 30, …
$ height <int> 62, 64, 64, 69, 67, 62, 62, 65, 62, 66, 68, 64, 63, 61, 63, …
$ weight <int> 100, 135, 115, 190, 125, 93, 178, 140, 125, 136, 120, 124, 1…
$ smoke <int> 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 0, 1, 1, 1, 0, 0, 1, 1, 0, 1, …16% of the variation in birth weight is explained by gestation. Higher values of R squared is preferred.
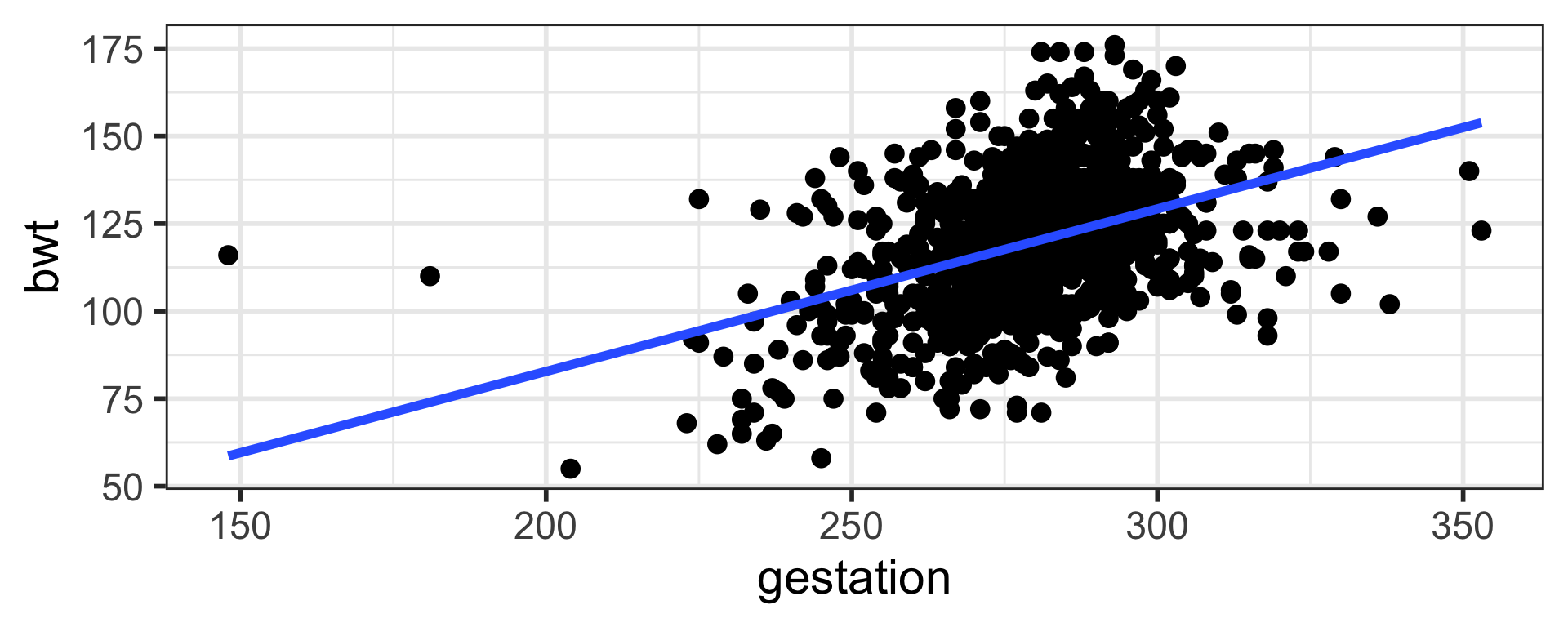
\(R\) and \(R^2\)
\(R^2\) = 0.1663449
\(b_1 = \frac{s_y}{s_x}R\)
\(R\) is the correlation coefficient
The Problem with Adding More Variables
- Every time you add a variable to a model, the regular
R-squaredwill always increase (or stay the same), even if the variable is useless. - This can trick us into building an overly complex model that is “memorizing” our sample data instead of learning the true underlying patterns. This is called overfitting (more on this later).
- An overfit model performs well on the data it was trained on, but fails when it sees new, unseen data.
A Better Metric: Adjusted R-Squared
- Adjusted R-squared penalizes the model for having extra variables that don’t contribute meaningfully.
- It only increases if the new variable improves the model more than would be expected by chance.
- Rule of Thumb: When comparing models with different numbers of variables, always prefer the one with the higher Adjusted R-squared.
Adjusted R-Squared
When comparing models with multiple predictors we rely on Adjusted R-squared.
Model Selection and Occam’s Razor
“Among competing hypotheses, the one with the fewest assumptions should be selected.”
How does this apply to our models?
- Simpler is Better: A model with fewer variables (a simpler model) is preferable to a complex model if they both have similar predictive power (similar Adjusted R-squared).
- Why?
- Avoids Overfitting: Simpler models are less likely to be overfit to the sample data and will generalize better to new data.
- Easier to Interpret: It’s much easier to understand the relationship between 3 variables and birth weight than between 10 variables and birth weight.
Occam’s Razor provides the philosophical foundation for why we use techniques like Adjusted R-squared and Train/Test splitting: to find the simplest model that does the best job.
# A tibble: 1,236 × 3
bwt pred resid
<int> <dbl> <dbl>
1 120 122. -1.79
2 113 121. -7.86
3 128 119. 8.53
4 123 NA NA
5 108 121. -12.9
6 136 123. 13.3
7 138 103. 34.8
8 132 104. 28.3
9 120 124. -4.11
10 143 129. 14.2
# ℹ 1,226 more rows# A tibble: 1,236 × 3
bwt pred resid
<int> <dbl> <dbl>
1 120 125. -4.80
2 113 125. -11.5
3 128 115. 13.3
4 123 NA NA
5 108 115. -7.47
6 136 125. 10.5
7 138 108. 30.4
8 132 107. 25.0
9 120 127. -6.81
10 143 124. 19.2
# ℹ 1,226 more rowsRoot Mean Square Error
\(RMSE = \sqrt{\frac{\Sigma_{i=1}^n(y_i-\hat y_i)^2}{n}}\)
Can we keep adding all the variables and try to get an EXCELLENT model fit?
Overfitting
We are fitting the model to sample data.
Our goal is to understand the population data.
If we make our model too perfect for our sample data, the model may not perform as well with other sample data from the population.
In this case we would be overfitting the data.
We can use model validation techniques.
Splitting the Data (Train vs. Test)
Exercises
8.12
9.4
9.10
